Update 2020: Anticovidian v.2 COVID-19: Hypothesis of the Lab Origin Versus a Zoonotic Event which can also be of a Lab Origin: https://zenodo.org/record/3988139
Author's Fernando Castro-Chavez Website, "Anthology of Works", My 2016 Scientific Research to the Journal of Biology and Nature:
Saved at the Archive dot Org as: http://liveweb.archive.org/web/20160208130349/http://fdocc.ucoz.com/index/vigesimal_mayan_twenty_aminoacids/0-140
And soon to be held also at the PubMed Database: http://www.ncbi.nlm.nih.gov/pubmed?term=%22Castro-Chavez%20F%22
Original research papers
Anatomical Mnemonics of the Genetic Code: A Functional Icosahedron and the Vigesimal System of the Maya to Represent the Twenty Proteinogenic Amino Acids.
Fernando Castro-Chavez [1]
Abstract
In programming and bioinformatics, the graphical interface is vital to describe and to abbreviate aspects and concepts of the physical world. The Mayan Culture developed the vigesimal system, a numerical system based on their count of fingers and toes. My objective is to equate the Mayan system and their numerical representation to the twenty amino acids according to size, except for the number one, represented by a dot, that here is given to cysteine, which acts as glue among peptides as one of its properties; in such a way, two vertical dots will be easily used to represent is related selenocysteine. The Mayan numerical system included the zero, represented by the Maya with an empty shell that here is used to represent the stop codons. On the other hand, the Chinese had a binary numerical system, similar to the binary comparisons of the three properties of Nucleotides within the double helix: H-Bonds, C-Rings and Tautomerism, called the I Ching which here is applied to the natural groups of amino acids that result of the 64-codons compared binary in their H-Bonds versus their C-Rings, used here to successfully represent the mature sequence of the glucagon amino acids. Additional anatomical tools for the mnemonics of the genetic code and of its amino acid groups are also presented as well as a functional icosahedron to represent them. Concluding, tools are presented for the visual analysis of proteins and peptide sequencing in bioinformatics and education to teach the genetic code and its resulting amino acids, plus their numerical systems.
Keyword, Vigesimal System, Yucatan, Chinese, I Ching, Yin/Yang, Maya, Glucagon
Introduction
Previously, it was demonstrated that the arrows of the Yin and Yang can be found contained in a 3-D fashion within a Cube where each of its walls is one of the six possible combinations of the three properties of the DNA Nucleotides: H-Bonds, C-Rings, and Tautomerism [1].
If we ask again: ‘What’s a code?’ the potential answer could be that “a code is like a dictionary, a set of correspondences between two different languages”, similar to a hidden message [2].
In the ‘intuitive’ “letter to number” code (the Roman numerals lacking zero), or “letter number” cipher, letters of an alphabet are replaced by numbers; for example, in English, we start with a-b-c…, making them to correspond to 1-2-3…, and we end with …x-y-z, being transformed, or ‘codified’ into …24-25-26.[2]
A variant of such “letter to number” code includes the zero (0) at the beginning (as the more advanced Arabic/Hindu numerical system did, also developed by the Maya, who had a vigesimal counting system based on the number of the human fingers and toes [3], a system that we are going to explore in this article and that is exemplified in Fig. 1.
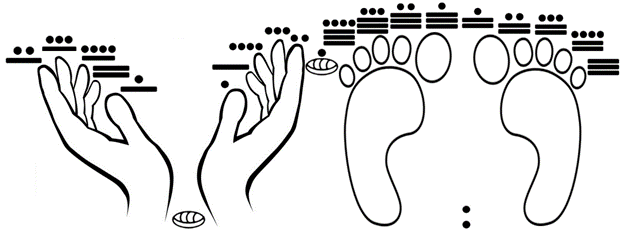
Figure 1. The Mayan vigesimal system based on the count of fingers and toes. The Maya also knew the zero (shown here between the wrists); for balance, the number 21 is also represented between the feet. Number one starts with the thumb of the right hand, proceeding orderly; then, number six starts with the thumb of the left hand, number eleven starts with the big toe of the right foot, and then number 16 starts with the big toe of the left foot, also proceeding orderly until the end.
So, in this ‘counter-intuitive’ system that includes the zero (0), we start with a-b-c…, which now will be transformed into 0-1-2…, and again ending with …x-y-z, which now will be numerically transformed into …23-24-25 [please notice this time the exclusion of the number 26 due to the inclusion of zero as an independent number, which also takes one space at the beginning]; we know that the inclusion of the zero was not only vital for the global development of arithmetic, and of commerce, but it’s also necessary for the Computer Sciences, and for programming codes in general [3].
For example, to say “I love you” with numbers in the “letter to number” code that includes the zero, the corresponding numbers are (using bold for capital letters, here the first letter, while adding an empty space to separate the words and a dash to concatenate letters within the same word):
8 11-14-21-4 24-14-20
The same word in the encryption lacking zero will be: 9 12-15-22-5 25-15-21. [3]
In the same way that ‘organically’ the Maya represented the numbers, we are going to review how to use our ten fingers in mnemonics to remember the most simpler ‘letter to number’ code; and by analogy, to also remember the genetic code. We are also going to use our four extremities, our two arms and our two legs, to remember the four foundational nucleotides of the genetic code: A, T, C, G.[4]
The “letter to number” hand-code ‘dictionary’ that we are going to use next does not include the zero as its starting point, we rather start with the number one, so a mnemonics letter to number code that could be used in this ‘letter number’ translation is proposed in Fig. 2.
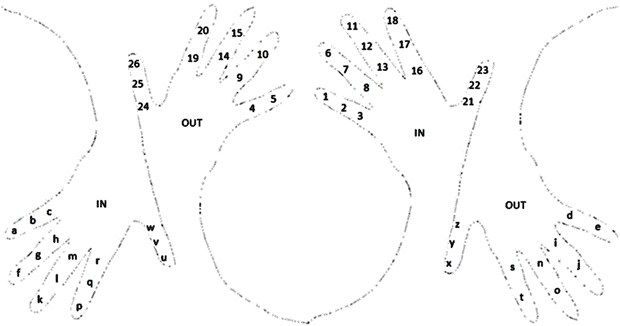
Figure 2. The use of the inner and outer side of each of our hands as mnemonics tools for the “letter to number” cypher translation.
Figure 2 shows that by using both of our hands as tools in mnemonics to remember the “letter to number” code, we use five letters or numbers to be memorized through each finger, except the thumbs that are used to memorize the last six of them. We start with the little finger (the ‘pinky’), then with the ring, the middle, the index, and thumb finger, from the inside of the hand, and in the inside from top to bottom as follows: distant phalanx, followed by the middle phalanx, and then by the proximal phalanx at the bottom of each finger, continuing with the next number, or letter, from the outside of the finger in the same relative position where we left as illustrated (in the outside from bottom to top); here, to identify which hand is which, as it was done the Mayan numerals in Fig. 1, just observe your own hands (right hand at the center, left at the extremes of Fig. 2).
In this proposal for mnemonics, the right hand contains the 24 numbers while the left hand holds the 24 letters of the English alphabet.
The skill, or mastery, to represent and to decipher hidden messages from one language into another, which most of the times is only known by two individuals, is known as ‘cryptography’ [2], and is highly used by detectives, spies, and their opponents; to be successful in this field, an acute memory is required, preferably to prevent from leaving traces of clues or evidences.
The case of the genetic code is quite different because, instead of wanting it to be only known ‘by the few’, as it happens in cryptography, the purpose of a good teacher is to teach the genetic code to as many persons as possible, while showing to them its intelligent design, and its complexities as simple as possible; however, currently most of the textbooks use only one page to represent the ‘square’ genetic code [5].
Four nucleotides in groups of three are called codons, or trinucleotides, and integrate our proteins, formed by the different arrangements that are possible between the twenty amino acids.
So, continuing with the physical metaphor, we will use our four extremities to represent the four nucleotides that integrate our DNA, the molecule located at the center of life: The whole of my right arm will represent Guanine (G) while my left arm will represent Cytosine (C); in the same way, my complete right leg will represent Adenine (A) while my left leg will represent Thymine (T, which in the RNA corresponds to Uracil, U).
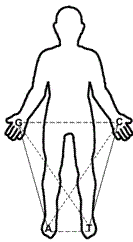
In Figure 3 we can see that when interlacing the hands, the triple hydrogen bonding that keeps together the double helix, is formed by the stronger step-ladder integrated by the components G and C (G-C); similarly, the weaker or double hydrogen bond is integrated by the weaker step-ladder A-T (or U in the RNA), which is anatomically exemplified here by putting our feet side-by-side.
Figure 3. Human extremities representing the four nucleotides G, C, A and T; solid lines indicate their identities according to their rings (purines: G and A; pyrimidines: C and T); the horizontal intermittent longer lines are the ones pairing nucleotides according to their hydrogen bonding (G-C and A-T), and finally, the intermittent diagonal shorter lines show the nucleotides that are able to experience a back and forth tautomerism (G to T and C to A).
Here, in Fig. 3, the right side (both right arm and right leg) are representing the bigger, double ringed purines, while the left side (both left arm and left leg) are representing the smaller one ringed pyrimidines.
The third property of nucleotides, their amino/imino tautomerism, is represented in this model (Fig. 3) by the opposite arms and legs, which means G is ‘tautomerizing’ to T (or to U) and C to A; making thus, easier to remember the three properties of nucleotides in the DNA: the horizontal hydrogen bonding, then their rings (vertical), and their tautomerism (crossed over, like in an “X” formation).
We can now go to the more complex use of our fingers in genetic code mnemonics, as presented in Fig. 4.
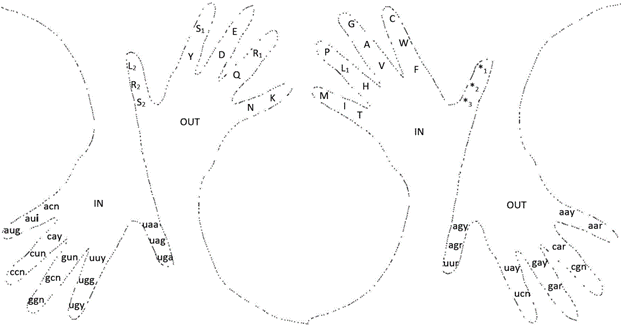
Figure 4. Use of both hands as tools in mnemonics to remember the genetic code and its codon-to-amino acid conversion; five codons, or amino-acids, are memorized by each finger, excepting the inner thumbs that are used as tools to memorize the three functional stop codons. We again start with the little finger, then with the ring, middle, index, and thumb, from the inside of the hand, top (distant phalanx), middle (middle phalanx), to bottom (proximal phalanx) of each finger, continuing the consecutive codon or amino acid in the outside of the finger in the same relative position where we left, as illustrated. To identify which hand is which, just look at your hands (right at the center, left at the extremes). Note: For the additional letters to the abbreviations: n: any nucleotide, r: purines, y: pyrimidines, i: A, C, T (or U). L1 and L2, R1 and R2, S1 and S2 represent the two different sets of codons that codify, respectively for L: Leu, R: Arg, and S: Ser.
The hand mnemonics shown in Fig. 4 integrate a code that can also provide a new pair of encryption codes for cryptography in the non-redundant forms of: 1) the number to amino acid, and 2) the letter to codon codification.
Finally, in this article I’m also going to provide a 3-D representation of the 20 proteinogenic amino acids, starting with representing them according to the Mayan numeration and to run the sequence glucagon amino acids through this table as a potential use for bioinformatics and educational tools.
Materials and Methods
The Mayan Vigesimal System used here was taken from:
The I Ching table used here was published elsewhere [1].
The amino acid sequence for the glucagon used here was taken from [15]:
H-S-Q-G-T-F-T-S-D-Y-S-K-Y-L-D-S-R-R-A-Q-D-F-V-Q-W-L-M-N-T
The amino acids selected for this study are those found on eukaryotes and not the ones in used in the biosynthesis of proteins in some methanogenic archaea and bacterium.
Results and Discussion
While the earlier focus of the Author has been in the codons [1, 6-11], this time the orientation of this work has been towards the amino acids themselves. The resulting correspondence between the Mayan vigesimal system and the amino acids according to their properties can be seen in Fig. 5.
 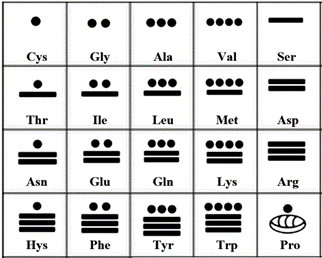 
Figure 5. Representation of amino acids according to the Mayan vigesimal system.
In the same way that the original symbols and arrangement of the I Ching were found to be adequate to represent the 64 codons of the Genetic Code, now the vigesimal system of the Maya is providing us with a complementary symbolic representation for the amino acids. If a coloring system were to be used, that will be the same as the one presented at [5], plus the addition of the Selenocysteine, Sec, U [12-13] whose RGB colors could be 255, 140 and 25; thus far, some 25 Sec proteins have been found in man [12].
The Mayan numbers present in Fig. 5 are independent of the numbers seen in Fig. 2.
The numbers start with the zero (an empty shell according to the Maya) for the stop function, then continue, for practical reasons, with Cys (C) as the number one, so that its related amino acid Selenocysteine (Sec), could be represented with a couple of vertical dots (being this the number 21); then, the successive amino acids in relation to their relative increase in size for the numbers from 2 to 19, will be: Gly (G), Ala (A), Val (V), Ser (S), Thr (T), Ile (I), Leu (L), Met (M), Asp (D), Asn (N), Glu (E), Gln (Q), Lys (K), Arg (R), Hys (H), Phe (F), Tyr (Y), Trp (W), followed by the bender of proteins: Pro (P).
If we focus now our attention in the groups of codons producing the 20 proteinogenic amino acids as they appear in the original representation of the I Ching, we obtain the results seen in Fig. 6, which in its distribution, contains the Mayan numbers and their corresponding amino acids according to the rows and groups of codons per amino acid that were shown elsewhere [1].
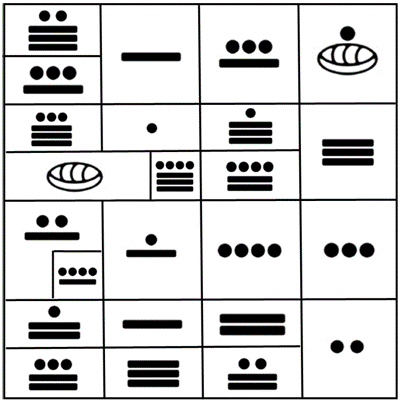
Figure 6. Groups of codons according to their amino acids from the original I Ching [1].
And now that we are considering vigesimal systems, if we include the icosahedron as the closest symmetry to represent the amino acids, in an ordering independent of the size of the amino acids as it was done to represent the amino acids using the Mayan system, we arrive at the results shown in Figure 7.
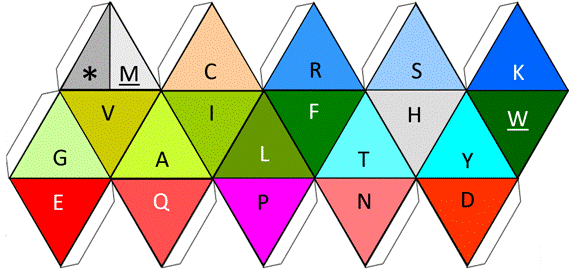
Figure 7. Functional icosahedron representing the twenty amino acids. The functional codons: start and finish, are represented as an on/off switch in the same cell, subdivided by a line (upper left corner).
The logic used to represent the 20 amino acids in Fig. 7, plus the functional stop by the use of a functional Icosahedron, is as follows: First, as its name declares and similar to the button turning the light on or off within our house, one of its cells includes the functional on/off switch able to work independently, with both functional options at the top (stop/Met, or */M), either as the point of entry or of exit. Followed by the Cys (C), which here acts as glue amongst peptides, while the basic hydrophilic amino acids, Arg (R) and Lys (K) are both also at the top surrounding the main phosphorilatable amino acid, Ser (S). Finally, and at the center of the functional icosahedron, we find the remaining two secondary phosphorilatable amino acids: Thr (T) and Tyr (Y), surrounding the central catalyzer/cutter of the enzymes: His (H). Then, at the center also, the rest are the hydrophobic amino acids, both the smallest: Gly (G) and Ala (A), and the biggest: Phe (F) and Trp (W), but also the rest of the intermediate ones: Val (V), Ile (I), and Leu (L), thus completing the design of the proteinogenic amino acids within the functional icosahedron. If a coloring system is to be desired, the one presented at [8] is the suggested option.
Others have used the angles of an icosahedron [14] and/or a combination of an icosahedron plus a tetrahedron [15], to represent all the amino acids, but not the triangular faces only of a functional icosahedron, as we do here, to represent the twenty amino acids plus their functional codons (the switch on/off).
Elsewhere, it was done a comparison of Human and of Neanderthal ‘analogous’ peptides [1] within faces of the genetic code cube; here, a practical example of the rotations of the functional icosahedron, and of its amino acids when located at the center of a sequence reader, is presented with glucagon as its example [16, 17] in Fig. 8; the design is unfolded for ease of view of the full trajectory that our sequence under consideration follows.
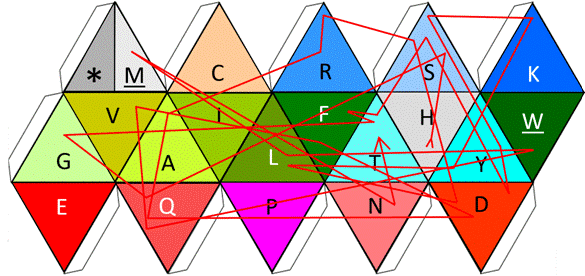
Figure 8. Representation of the reading of the amino acids of glucagon by the functional icosahedron.
In Fig. 9 we can see both the static and the dynamic use of the functional amino acid configuration proposed here for bioinformatics and for educational applications; we are able to see, both the folded structure of the functional icosahedron amino acid (left side) as the center engine of a bioinformatics program before its rotation starts; a sequence can be fed to be read, such as the sequence of glucagon used here as example to see a portion of its rotational directions through its on/off (start/end) face.
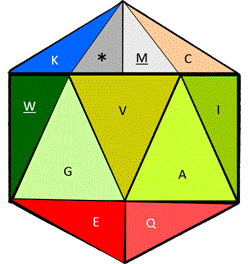 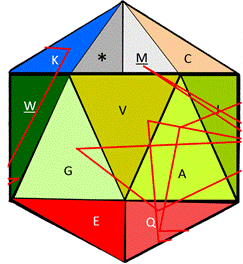
Figure 9. Folded representation of the functional amino acid icosahedron in its static start position, without any sequence to read (left side), and the lines of the rotational directions according to amino acids (there, one amino acid is touched three times (Q), wile 6 amino acids are touched only once, in the current face in the following visible order: G, K, A, V, W, and M) corresponding to the mature version of the glucagon protein.
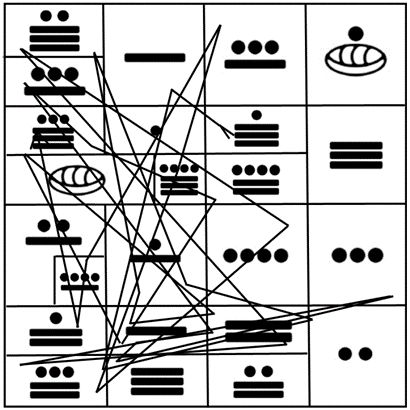
Figure 10 shows a practical example with the sequence of glucagon [16, 17] run through the first face of the cube represented in Fig. 6, similar to the initial comparison of analogous peptide sequences between Humans and Neanderthals shown elsewhere [1].
Figure 10. Example of the human sequence of glucagon as run within the first face of the cubic representation of the genetic code introduced elsewhere [1], with the vigesimal number of the Maya indicating their corresponding amino acids.
Finally, if we now include the amino acids within the vigesimal numerical system of the Maya, we obtain the results displayed in Fig. 10, where the twenty fingers and toes are used to represent the main twenty proteinogenic amino acids.
Conclusion
It was, not only possible to represent a correspondence between the codons and their amino acids by using the inner and outer part of both hands, but also the amino acids were correlated to the vigesimal system of the Maya, and then each finger and toe were used to represent each one of the amino acids.
At the end, a practical application was shown where the amino acid sequence of glucagon was run through one resulting table of the binary comparison of H-Bonds in axis X versus C-Rings in axis Y, with the Mayan numbers occupying each one of the cells representing groups of codons per their resulting amino acid.
When comparing the detailed representation of each codon as presented in Fig. 4 with the more encompassing one of grouping them within their resulting amino acids and functions, shown by either the icosahedron (Fig. 7) or the Mayan numerals (Fig. 5), it’s easy to realize that the compressed representation of codons shown by these last ones is easier to memorize and to handle programmatically than the first and most extended one.
It was also possible to represent the 20 amino acids within a regular functional icosahedron, with one specific modification: The functional codons (start and stop) should be shared within the same cell while operating independently, such as in the on/off switch for the light still currently present in almost every house.
This article was mostly focused on the graphical representation of the twenty amino acids by using a vigesimal system, the Mayan, which is less known than the 64-esimal system of the I Ching; as a reviewer wisely noticed, further research is needed, both for the graphical design of tools to compare sequences of amino acids and of genes, as well as the practical advantage for the students when taught using these two different representations based on the ancient ways to define and to represent numbers, with the conventional way of teaching.
As a bonus, a hands-on representation of my previous research [8] using the OpenSCAD:
Acknowledgments
Ana Baleva suggested the use of the icosahedrons to represent the twenty amino acids. I wish to thank Jose Francisco Norambuena Michea for accepting me in his Graphical Design team in Chile (Norambuenagrafic Polarizados), and to Barbara Larreategui, the first one to whom I took the reciprocal initiative to say: 8 11-14-21-4 24-14-20 (see above); this manuscript was partly supported by the NIH grant T32 HL-07812.
References
1. Castro-Chavez F: File compression and expansion of the genetic code by the use of the Yin/Yang directions to find its sphered cube. J Biodivers Bioprospect Dev 2014, 1:112. URL: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4203674
2. Bishop D: Introduction to cryptography with Java applets (pp. 16-17). Jones & Bartlett Learning 2003, 370 pp. http://books.google.com/books?id=yxPnt4S3mFMC
3. Sharer RJ: The ancient Maya. Stanford University Press 2006, 931 pp.
4. Ifrah G: A universal history of computing: from the abacus to the quantum computer. John Wiley 2001, 410 pp.
6. Castro-Chavez F: The rules of variation: amino acid exchange according to the rotating circular genetic code. J Theor Biol 2010, 264:711-21. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3130497
7. Castro-Chavez F: The rules of variation expanded, implications for the research on compatible genomics. Biosemiotics 2012, 5:121-145.
8. Castro-Chavez F: Defragged binary I Ching genetic code chromosomes compared to Nirenberg’s and transformed into rotating 2D circles and squares and into a 3D 100% symmetrical tetrahedron coupled to a functional one to discern start from non-start methionines through a stella octangula. JPSCB 2012, 1:1-24. http://www.hoajonline.com/jpscb/2050-2273/1/3
9. Castro-Chavez F: A tetrahedral representation of the genetic code emphasizing aspects of symmetry. Bio-Complexity 2012, 2:1-6. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3445437
10. Castro-Chavez F. The quantum workings of the rotating 64-grid Genetic Code. Neuro Quantology 2011; 9:728-746. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3271378
11. Castro-Chavez F. Most used codons per amino acid and per genome in the code of man compared to other organisms according to the rotating circular genetic code. NeuroQuantology 2011; 9:747-766. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3445418
12. Gonzalez-Flores JN, Shetty SP, Dubey A, Copeland PR. The molecular biology of selenocysteine. Biomol Concepts 2013; 4:349-65.
13. Kryukov GV, Castellano S, Novoselov SV, Lobanov AV, Zehtab O, Guigo R, Gladyshev VN. Characterization of mammalian selenoproteomes. Science 2003; 300:1439–1443.
14. Karasev VA, Luchinin VV, Stefanov VE. Symmetry and spatial structure of the canonical set of amino acids, Proceedings of the BGRS, 2004, pp. 278-281.
15. Pitkanen M. Pythagoras, music, sacred geometry, and genetic code, Self-published by Pitkanen, 2014, 21 pp.
16. Bromer WW, Sinn LG, Staub A, Behrens OK. The amino acid sequence of glucagon. Diabetes 1957; 6:234-8.
17. Dods RF. Understanding diabetes: A biochemical perspective. Wiley 2013; 426 p.
[1] Department of Medicine, Atherosclerosis and Vascular Medicine Section, Baylor College of Medicine, Houston, TX, USA. Phone: +1 713 798 4177. Fax: 713-798-4121. e-mail: fdocc@yahoo.com
[3] As per the suggestion of a reviewer, the most common sentence in programming: “Hello World” can be represented by the encryption system having the zero as: 9-6-13-13-16 24-16-19-12-5 and by the one lacking zero as: 8-5-12-12-15 23-15-18-11-4.
[4] Abbr.: Amino acids: A: alanine (Ala), V: valine (Val), I: isoleucine (Ile), L: leucine (Leu), M: methionine (Met), F: phenylalanine (Phe), W: tryptophan (Trp), D: aspartic acid (Asp), N: asparagine (Asn), E: glutamic acid (Glu), Q: glutamine (Gln), R: arginine (Arg), K: lysine (Lys), S: serine (Ser), T: threonine (Thr), G: glycine (Gly), P: proline (Pro), H: histidine (His), C: cysteine (Cys), Y: tyrosine (Tyr). Nucleotides (nt): U: uracil, C: cytosine, A: adenine, G: guanine, T: thymine. n: any nt, r: purines, y: pyrimidines, i: A, C, T (or U).
|









